The ABCD_BH034 antibody recognizes the phosphorylated 53BP1 peptide by ELISA
DOI:
https://doi.org/10.24450/journals/abrep.2026.e2511Abstract
The engineered Forkhead-associated Domain (FHA1), located at the N-terminus of the RAD53 protein of Saccharomyces cerevisiae, was shown in previous studies to specifically recognize the phosphorylated peptide of the p53-binding protein (53BP1). When the engineered FHA1 domain was expressed as a recombinant rabbit Fc-fusion protein, it still recognized the phosphorylated 53BP1 peptide in an ELISA. FHA1-Fc fusion proteins can thus be used as specific antibodies to identify the phosphorylation status of various proteins.
Introduction
More than two-thirds of the 21,000 human proteins are phosphorylated, yielding more than 200,000 known phosphorylation sites (www.phosphosite.org/). Most of the phosphorylation events occur on serine residues (86.4%), followed by threonine (11.8%) and tyrosine (1.8%). Protein phosphorylation plays critical roles in signal transduction, protein localization, protein-protein interactions, transcriptional regulation, and enzyme activation, while aberrant phosphorylation events are associated with numerous diseases, including cancer, Alzheimer’s, and other disorders. Thus, accurate monitoring of protein phosphorylation is of great importance (Ardito et al., 2017). The RAD53 protein of Saccharomyces cerevisiae plays a critical role in arresting cell division in response to DNA damage by regulating proteins such as Asf1, which is involved in chromatin assembly. Rad53 regulates Asf1 through direct interaction, mediated by the phosphothreonine recognizing Forkhead-associated domain (FHA1) located at the N-terminus of Rad53 (Emili et al., 2001). In our previous work (Thota et al., 2024), we screened a phage library displaying an engineered FHA1 domain and identified binders to a variety of peptides phosphorylated at central threonine residues. We have termed these engineered FHA domains "phosphothreonine binding domains" (pTBDs). We validated a pTBD specific for the 53BP1 peptide phosphorylated at threonine (53BP1-pT543; 536-IDEDGENpTQIEDTEP-551). Notably, this pTBD distinguished between phosphothreonine and phosphoserine, which differ by only a single methyl group (Thota et al., 2024). The human TP53-binding protein 1 (53BP1; Uniprot #Q12888) is a multifunctional protein involved in several key cellular processes. It plays an important role in repairing double-stranded DNA damage by non-homologous end joining (NHEJ) by recruiting several factors during DNA damage (Lei et al., 2022). In addition, 53BP1 regulates the tumor suppressor protein p53 and maintains replication fork stability during cell division. 53BP1 is phosphorylated at T543 in an ATM/ATR-dependent manner during DNA damage (Stokes et al., 2007). Thus, phosphorylated 53BP1 at T543 is an important target for monitoring. In this study, we demonstrated that the recombinant fusion of the 53BP1-pTBD to rabbit Fc is functional and specifically binds to 53BP1pT peptide, as determined by ELISA.
Materials & Methods
Antigen binder: 53BP1-pTBD (ABCD_BH034, ABCD nomenclature, http://web.expasy.org/abcd/) was produced by ABCD Antibodies (https://abcd-antibodies.com) as engineered 53BP1-pTBD fused to a rabbit IgG Fc. HEK293 suspension cells growing in HEK TF medium (Xell #861-0001, Sartorius), supplemented with 0.1% Pluronic F68 (Sigma #P1300) were transiently transfected with the vector coding for the 53BP1-pTBD-Fc (ABCD_BH034-Fc) fusion protein. Supernatants were collected after 4 days.
Antigen: Peptides derived from human mechanistic target of rapamycin (mTOR) NKSRTRpTDSYSAGQ (mTOR-pT), CHK2 checkpoint homolog (Chk2) SSLETVSpTQELYSIP (Chk2-pT), tumor suppressor p53-binding protein (53BP1) IDEDGENpTQIEDTEP (53BP1-pT), and ataxia telangiectasia and Rad3-related protein (ATR) LCFPENEpTPPEGKNM (ATR-pT) were synthesized with an N-terminal biotin and a phosphorylated threonine residue by LifeTein (Somerset, NJ, USA). Phosphoserine and non-phosphorylated variants of 53BP1 peptide, IDEDGENpSQIEDTEP (53BP1-pS) and IDEDGENTQIEDTEP (53BP1-T), were also included as targets in the ELISA.
Protocol: Biotinylated phosphorylated peptides ATR-pT, mTOR-pT, Chk2-pT, 53BP1-pT, and 53BP1-pS were captured on a NeutrAvidin-coated high-binding microtiter plate at a concentration of 100 nM. Non-specific binding sites in the wells were blocked with 2% milk in PBS for 1h. After blocking, the wells were washed once with PBS-0.05% Tween 20 (PBST), and the ABCD_BH034 was added to the wells at 450 ng/µL concentration. After a 1h incubation, the plate was washed three times with PBST. Horseradish peroxidase (HRP)-conjugated α-Rabbit (Jackson ImmunoResearch Laboratories, Inc., Cat#111-035-003) antibodies were diluted 1:5000 in PBST, added to the wells, and incubated for 1 h. The plate was then washed three additional times with PBST. The HRP substrate 3,3’,5,5’ tetramethylbenzidine (TMB) (Thermo Scientific, Cat#34022) was added to the wells and allowed to develop for 1-2 minutes. The reaction was quenched by adding 1N HCl, and the absorbance was measured at 450 nm in a BioTek Synergy HTX Multimode Reader.
Results & Discussion
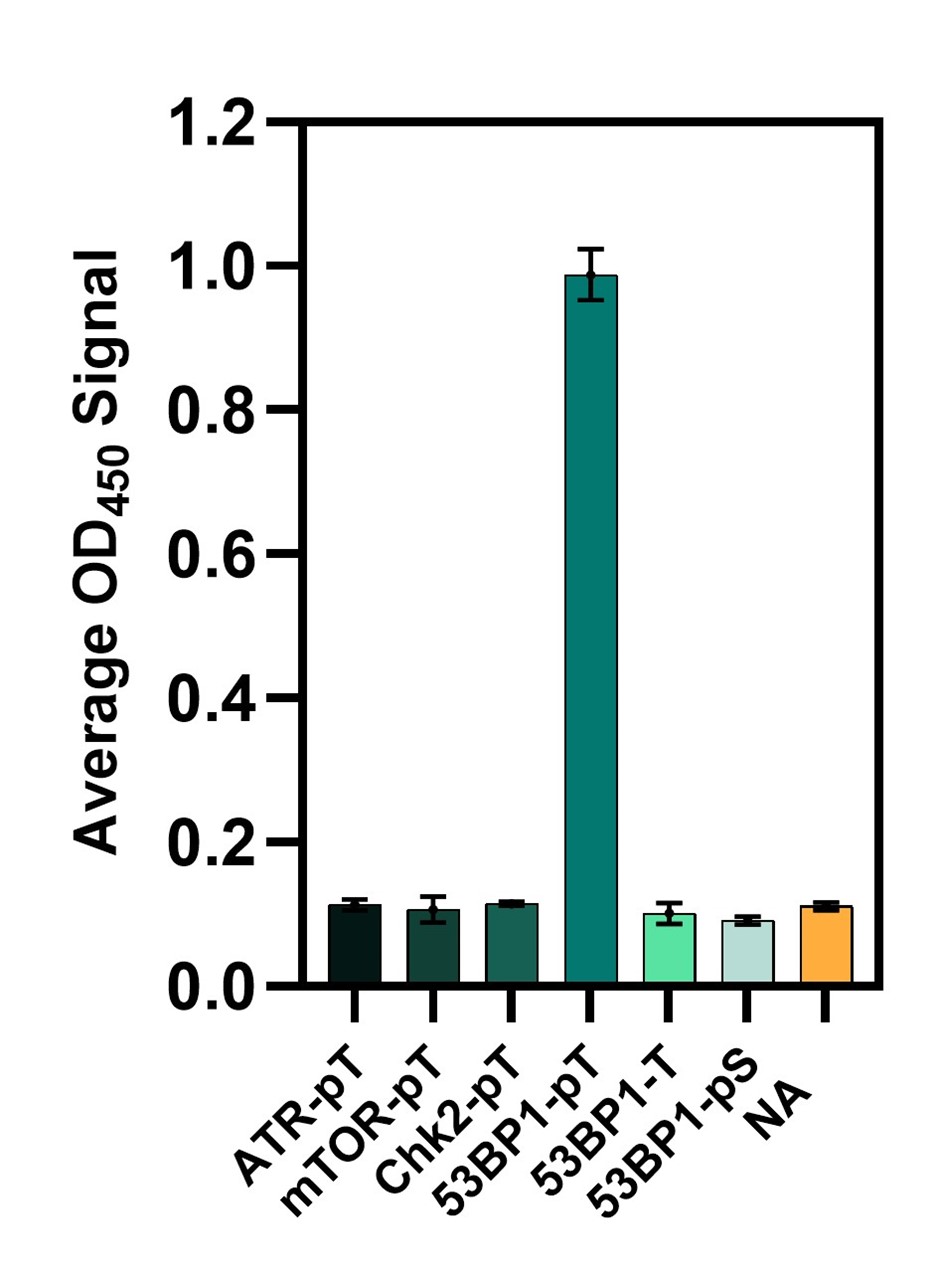
53BP1-pTBD (ABCD_BH034) was produced as a rabbit Fc fusion (ABCD_BH034-Fc). When evaluated by ELISA (Fig. 1), this antibody-like molecule specifically recognized the 53BP1 pT peptide. Notably, ABCD_BH034-Fc did not bind to a 53BP1-derived peptide containing a phosphorylated serine instead of a threonine, despite the two peptides differing by only a single methyl group. Furthermore, ABCD_BH034-Fc showed no background binding to other threonine-phosphorylated peptides. The detection of post-translational modifications in diagnostic assays remains challenging. The development of ABCD_BH034, therefore, provides a foundation for generating antibody-like proteins that recognize post-translational modifications with improved specificity and accuracy.
Figure 1. Binding of ABCD_BH034-Fc fusion protein to a collection of phosphopeptides. ABCD_BH034 does not bind to peptide 53BP1-pS (phosphoserine) or other phosphothreonine peptides and NeutrAvidin (NA), as detected by ELISA. The standard deviation of triplicate measurements is represented by error bars.
Conflict of interest
Srinivas Thota and Brian Kay are employees of Tango Biosciences, a contract research organization that discovers peptide ligands and recombinant affinity reagents for clients. Brian Kay is also a co-founder and shareholder. These affiliations do not alter the authors’ adherence to journal policies.
Data Availability Statement
The data that support the findings of this study are available from the corresponding author upon reasonable request.
References
Ardito, F., Giuliani, M., Perrone, D., Troiano, G., & Lo Muzio, L. (2017). The crucial role of protein phosphorylation in cell signaling and its use as targeted therapy (Review). International Journal of Molecular Medicine, 40(2),271–280. https://doi.org/10.3892/ijmm.2017.3036
Emili, A., Schieltz, D. M., Yates, J. R., & Hartwell, L. H. (2001). Dynamic Interaction of DNA Damage Checkpoint Protein Rad53 with Chromatin Assembly Factor Asf1. Molecular Cell, 7(1), 13–20. https://doi.org/10.1016/s1097-2765(01)00150-2
Lei, T., Du, S., Peng, Z., & Chen, L. (2022). Multifaceted regulation and functions of 53BP1 in NHEJ mediated DSB repair (Review). International journal of molecular medicine, 50(1), 90. https://doi.org/10.3892/ijmm.2022.5145
Stokes, M. P., Rush, J., Macneill, J., Ren, J. M., Sprott, K., Nardone, J., Yang, V., Beausoleil, S. A., Gygi, S. P., Livingstone, M., Zhang, H., Polakiewicz, R. D., & Comb, M. J. (2007). Profiling of UV-induced ATM/ATR signaling pathways. Proceedings of the National Academy of Sciences of the United States of America, 104(50), 19855–19860. https://doi.org/10.1073/pnas.0707579104
Thota, S. S., Allen, G. L., Grahn, A. K., & Kay, B. K. (2024). Engineered FHA domains can bind to a variety of Phosphothreonine-containing peptides. Protein Engineering, Design, and Selection, 37, gzae014. https://doi.org/10.1093/protein/gzae014.
Downloads
Published
Section
How to Cite
License
Some rights reserved 2026 Srinivas S. Thota, Brian K. Kay

This work is licensed under a Creative Commons Attribution 4.0 International License.



